Multiple File Datasets¶
Introduction¶
Traditional Galaxy workflows require a fixed number of input files and may produce a large number of intermediate files per input. Additionally, Galaxy renames datasets with each step, making tracking samples or fractions across long workflows very difficult. These issues together make Galaxy a problematic platform for workflows and application areas that require dealing with a large number of files. The prevalence of fractionated samples in mass spectrometry makes proteomics such an application area.
Galaxy-P however utilizes an extension to the core Galaxy framework called “Multiple File Datasets” designed to address these shortcomings in Galaxy. In simple terms, Galaxy-P can group many similar files into a single dataset (called a multiple file dataset). Normal Galaxy tools can then use these datasets and produce multiple file datasets of their own, operating on each item in parallel. In addition to keeping the number of datasets in the Galaxy managable, this allows for the creation of workflows that can operate over any number of files and the individual files in the multiple file dataset are given consistent, trackable names across such complex workflows making sample tarcking trivial.
Creating a Multiple File Dataset¶
Once you have an initial multiple file dataset, most tools when operating on these datasets will in return produce multiple file dataset outputs. So for most proteomic workflows, the first step is simply to create a multiple file RAW dataset or a multiple file peak list (e.g. mzML or mgf) dataset.
There are several ways to do this.
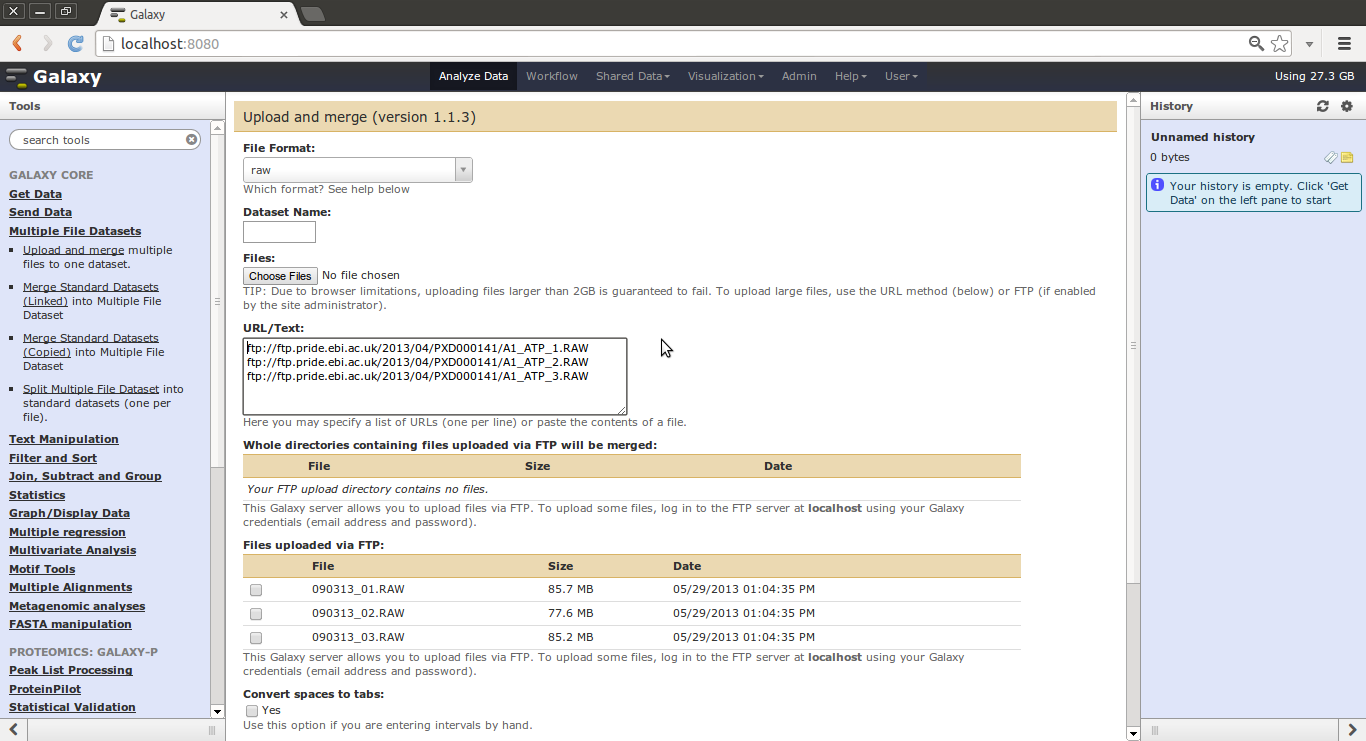
One can use the “Multiple File Datasets” -> “Upload and Merge” tool along to create a multiple file datasets from files or a directory of files uploaded via FTP.
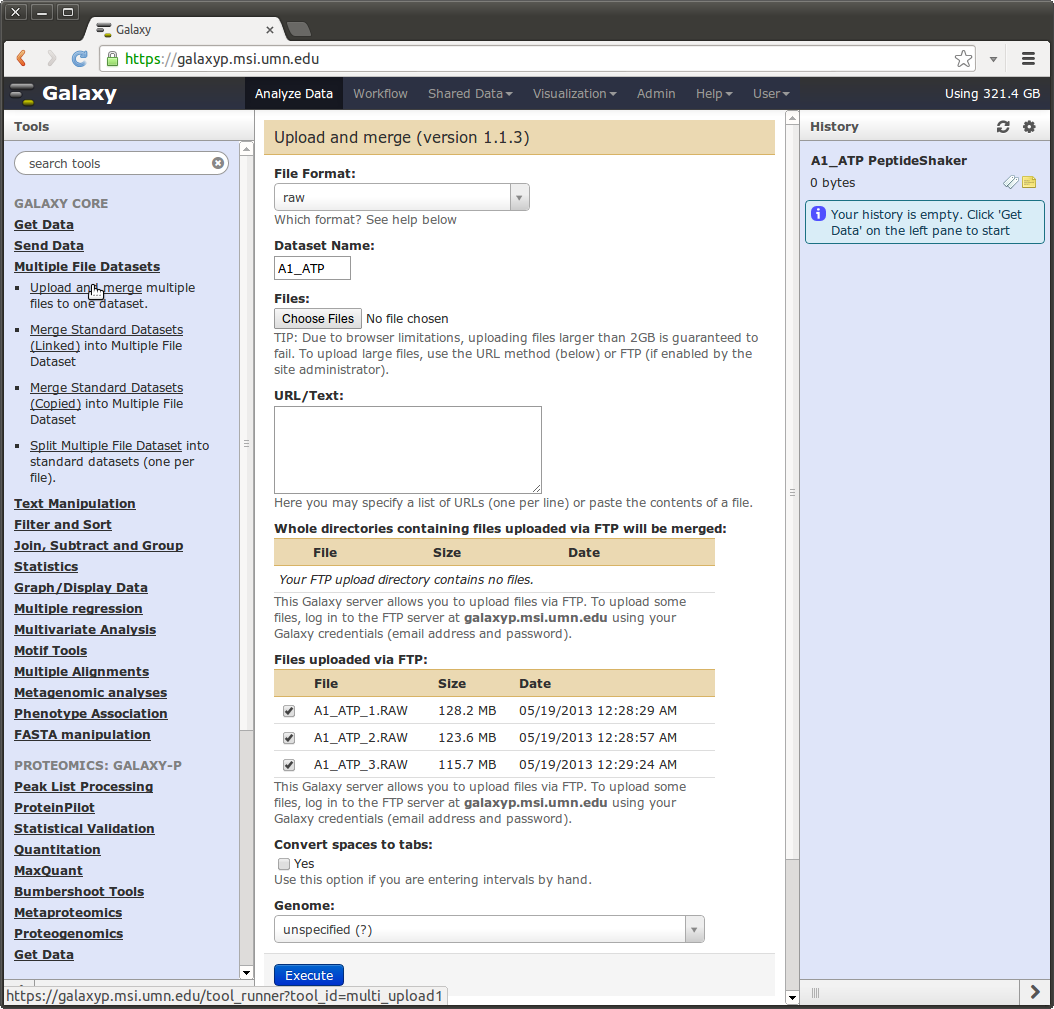
One can use the “Multiple File Datasets” -> “Upload and Merge” tool and simply click the “Choose Files” button and select multiple file for upload.
One can use the “Multiple File Datasets” -> “Upload and Merge” tool and paste in multiple URLs to have Galaxy download these files and create a multiple file dataset.